A genome-resolved understanding of rice-paddy microbial
communities and methane dynamics
PC Bethany Kolody
Rice is an important staple food, accounting for 19% of global calories and 13% of global protein in 2009. As the world population continues to grow, rice production will need to expand even further to meet rising demands. Unfortunately, the cultivation of rice is also a major green house gas contributor. It accounts for about 10% of anthropogenic methane emissions. Rice production emits methane because rice paddies are flooded. Flooding prevents the emergence of weeds, but also creates an anoxic soil environment that forces carbon (e.g. rice straw, root exudates) to be broken down anaerobically, eventually leading to methane as an end product.
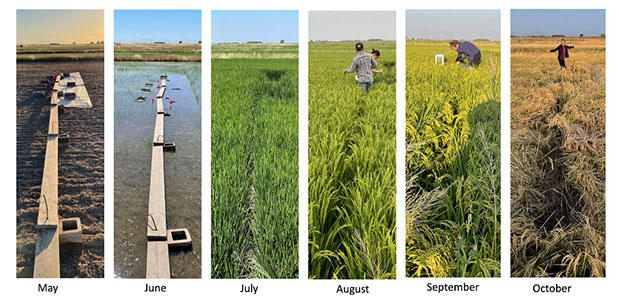
While significant strides have been made in understanding the microbiology responsible for methane production in rice fields, most in-situ studies rely on marker-genes to identify the key players. Here, in a pilot experiment, we sampled a paddy at the Rice Experiment Station in Biggs, CA over the course of the 2021 growing season pairing metagenomics and metatranscriptomics to gas fluxes over time. Our goal is to use genome-resolved metagenomics to elucidate the metabolic potential of the key players in the system, and metatranscriptomics to understand how environmental conditions drive methanogenesis and methanotrophy. The field-based experiments will continue in the next two years.
In a second project in collaboration with Prof. Pam Ronald, UC Davis, we will probe how rice genetic variants differ from wildtype rice in terms of their root-associated microbial communities and potential for greenhouse gas emissions. The initial phase is a greenhouse-based study but experiments in the following two years will likely transition into field studies in rice paddies. Ultimately, we hope that this will yield insights that will inform rice growing best practices that mitigate methane emissions for a
greener future.
Methane-oxidizing archaea and their extraordinary Borg extrachromosomal elements
Working in wetland soils we discovered surprising large, linear, peculiar genetic elements that we infer associate directly with the anaerobic methane oxidizing Methanoperedens archaea. Notably, these elements encode many genes that are directly implicated in methane oxidation metabolism. We will
continue to investigate the abundance, distribution, evolution and potential biogeochemical importance of this association and hope to learn how to harness this partnership to reduce methane emissions from anaerobic, carbon-rich environments.